@InProceedings{Alfasly_2024_CVPR,
author = {Alfasly, Saghir and Shafique, Abubakr and Nejat, Peyman and Khan, Jibran and Alsaafin, Areej and Alabtah, Ghazal and Tizhoosh, H.R.},
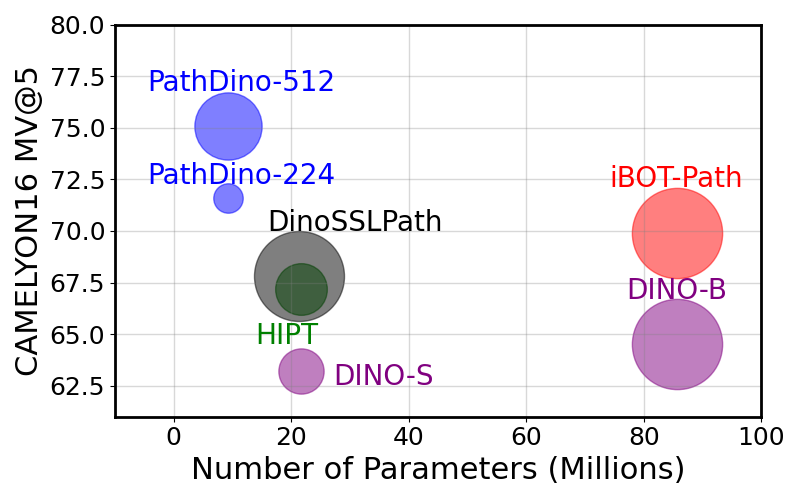
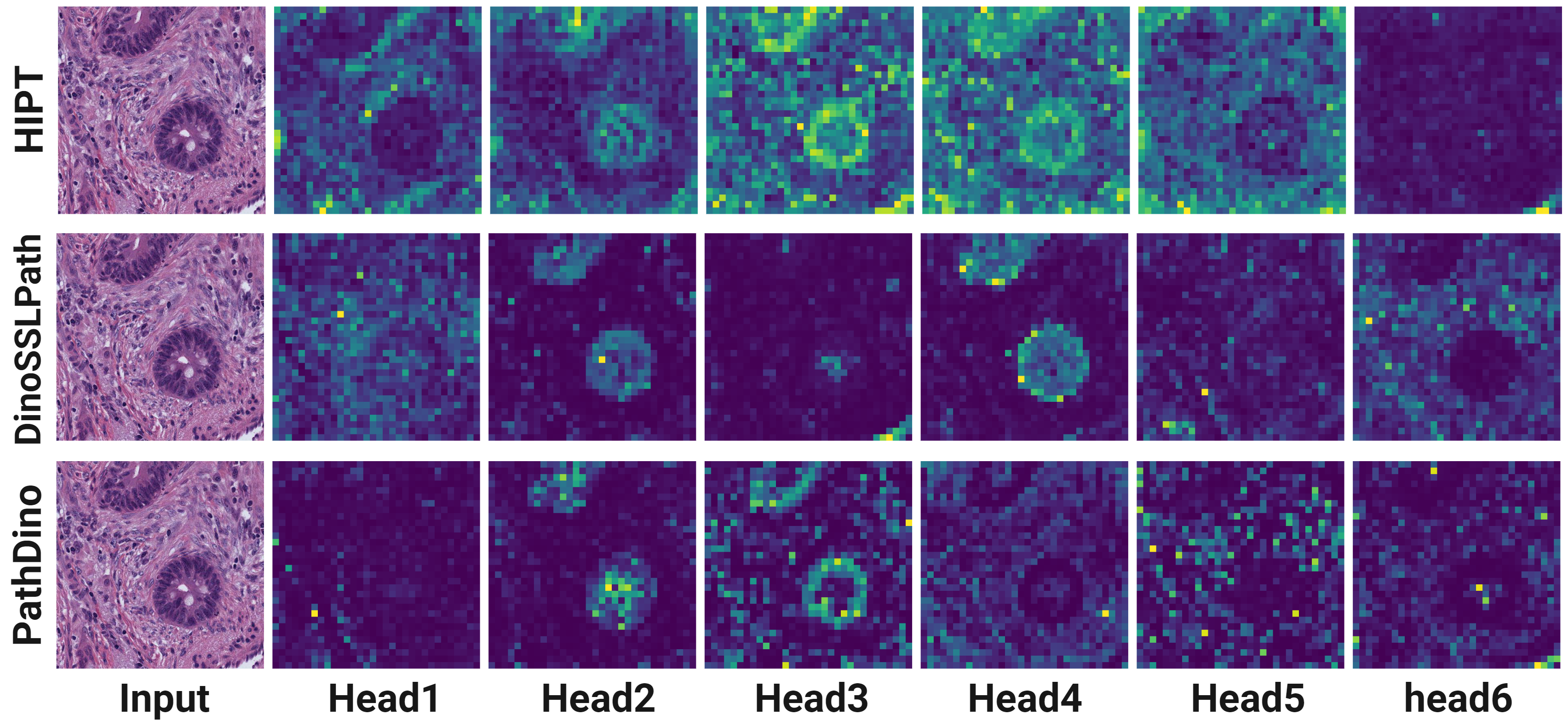
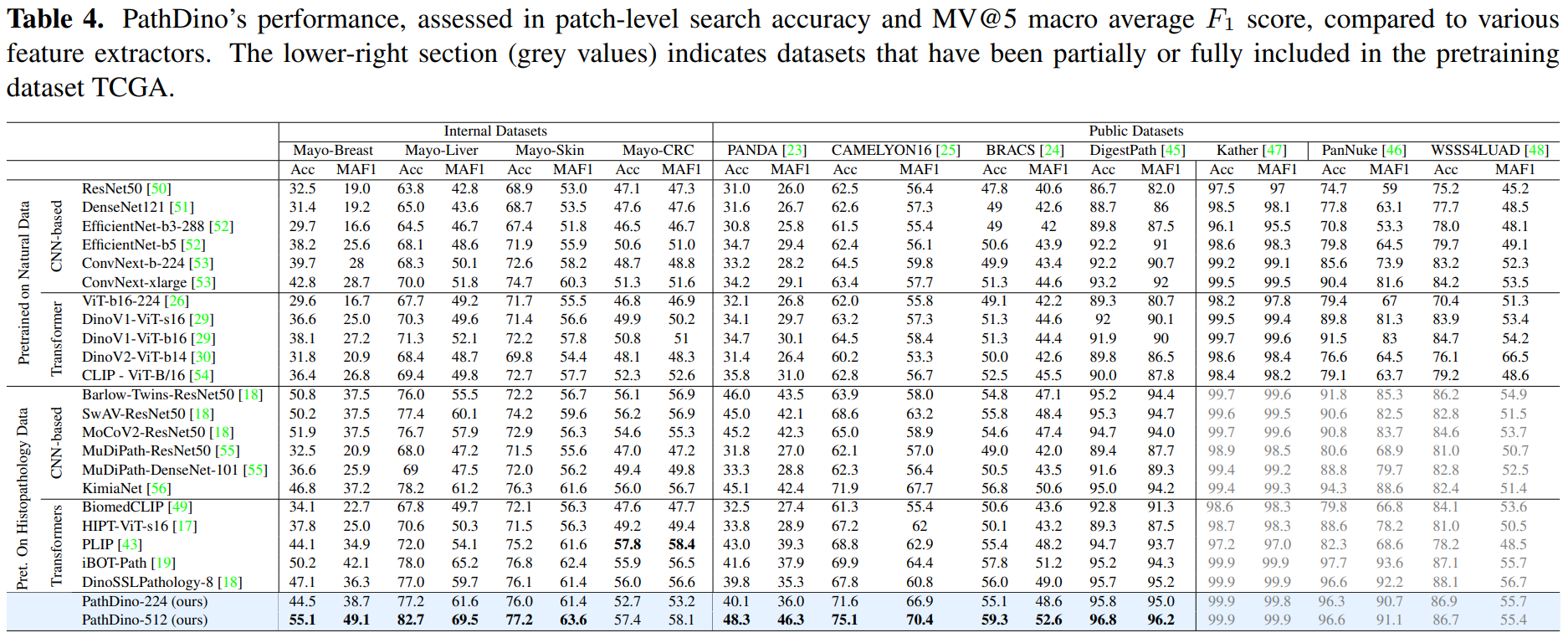
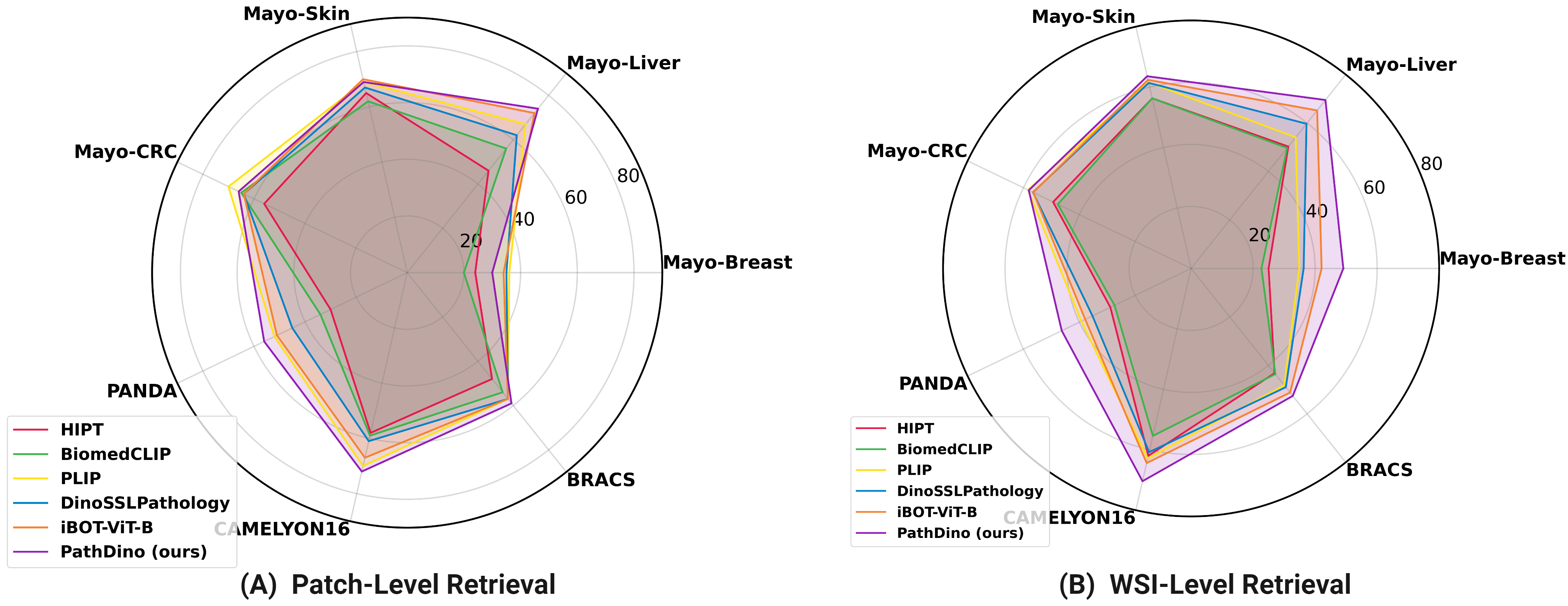
title = {Rotation-Agnostic Image Representation Learning for Digital Pathology},
booktitle = {Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR)},
month = {June},
year = {2024},
pages = {11683-11693}
}
KIMIA Lab, Department of Artificial Intelligence & Informatics, Mayo Clinic, Rochester, MN, USA
This web page template is borrowed from here.